AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
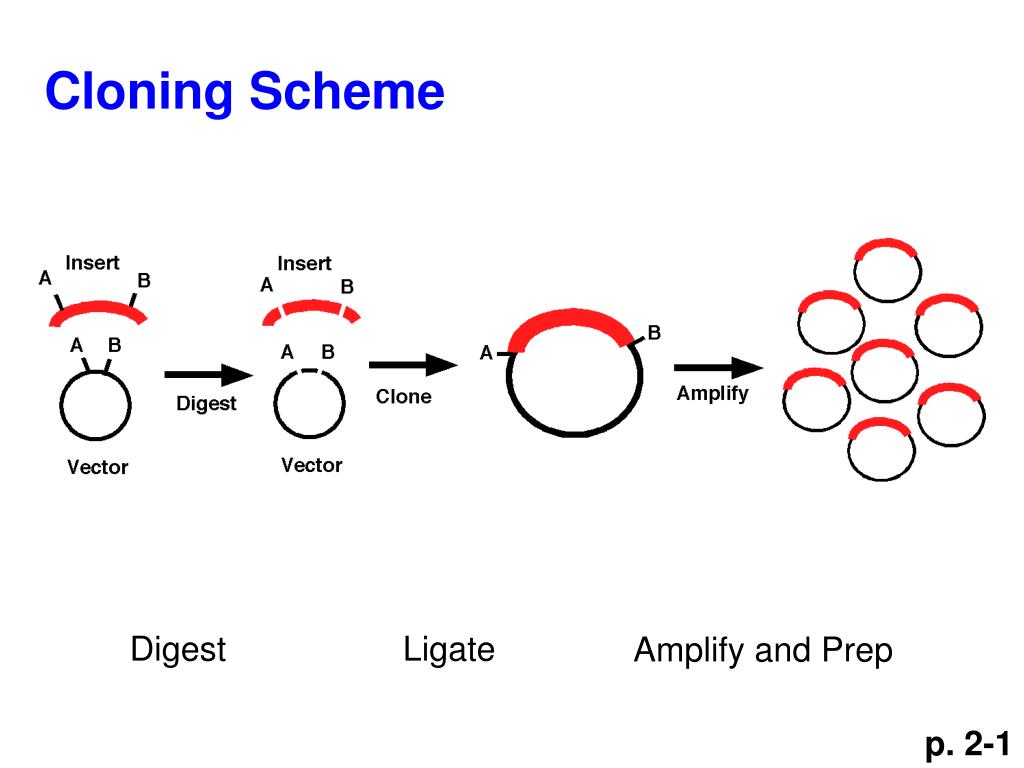
Taq polymerase preferentially adds a single 3'-A to blunt-ended, double-stranded DNA via this terminal transferase-like activity (13, 14). It has been observed that many DNA polymerases, including Thermus aquaticus (Taq) DNA polymerase, are capable of adding an additional non- template directed nucleotide to the 3'-ends of a blunt-ended DNA fragment. Several methods have now been developed for the cloning of PCR-amplified DNA molecules, including cohesive-end cloning by the introduction of restriction sites at the 5'-ends of PCR primers (7), blunt- end cloning (8, 9), ligation- independent cloning (3-6), and TA cloning (10, 11, 12). Although the development of PCR made the rapid amplification of DNA fragments generally quite simple, the cloning of PCR products was often found to be quite difficult using early technology. Both of the ligase-dependent methods require multiple purification and/or enzymatic modification steps. The other, less efficient approach is blunt-end ligation in which blunt-ended DNA fragments are ligated to a linearized blunt-ended plasmid. However, for every restriction fragment having distinct cohesive ends, one linearized plasmid vector compatible with that (those) particular restriction site(s) has to be prepared. This approach is the most efficient of the two. One approach is cohesive-end ligation, in which cohesive ends on the insert and plasmid are generated by digestion with appropriate restriction enzymes, and the complementary ends of the plasmid and insert are then joined by DNA ligase. There are two subclasses of ligase-dependent methods employed for the covalent joining of a DNA insert to a plasmid vector. This issue was easily resolved conducting an additional Although, the In-Fusion ® cloning method can be used for fusing several fragments in one reaction, in our experience, increasing the number of fragments leads to reduced number of positive clones. The In-Fusion ® enzyme uses a 15-bp overlap between to-be-fused fragments, which can be easily incorporated into primers that are used for PCR amplification. Using this plasmid as a template, we have developed a robust clon- ing method for direct cloning of gRNAs in a plant expres- sion vector using the ligation independent In-Fusion ® cloning system. We developed a unique template plasmid, pEN-Chimera- ccdB carrying the ccdB gene, which virtually eliminated any background plasmids thus greatly accelerating the cloning process for making multiple gRNAs. Taking advantage of this property, multiple cloning methods such as the Gateway ® cloning methods have been developed for reducing background colonies. coli cells that do not carry the antitoxin ccdA gene. The ccdB gene codes for a toxin, which is part of toxin-antitoxin system that inactivates DNA-topoisomerase II complexes thus kill- ing host E. The system developed here utilizes negative selection based on the ccdB gene. This requires screening many colonies to identify positive clones consuming resources and time. Ligation-dependent methods are also prone to getting a larger number of background colonies resulting from the undigested parent vector. Current methods for cloning the target-specific protospacer sequence with an RNA pol III-controlled promoter and the scaffold of sgRNA are time consuming and involve multiple expensive clon- ing steps utilizing type IIS restriction enzymes. The gRNA construct is then usually cloned in a plant expression vector, which carries the Cas9 gene on the same T-DNA cassette. The protospacer for each target has to be cloned between an RNA pol III-controlled promoter and the scaffold of sgRNA. The type II CRISPR/Cas9 system consists of a fixed core RNA-guided DNA nuclease, Cas9, and a chimeric gRNA consisting of a non-variable RNA scaffold derived from the crRNA and tracrRNA and a target-specific 20-nucleotide protospacer RNA, which is engineered specifically for a genomic region to be edited. The CRISPR/Cas9 system has rapidly become a method of choice for genome editing in basic and applied research in various organisms.
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
